forked from ThomasCarroll/Bioconductor_Introduction
-
Notifications
You must be signed in to change notification settings - Fork 2
Log2 transformed bigWig not opening in IGV/exporting correctly #11
Copy link
Copy link
Open
Labels
Description
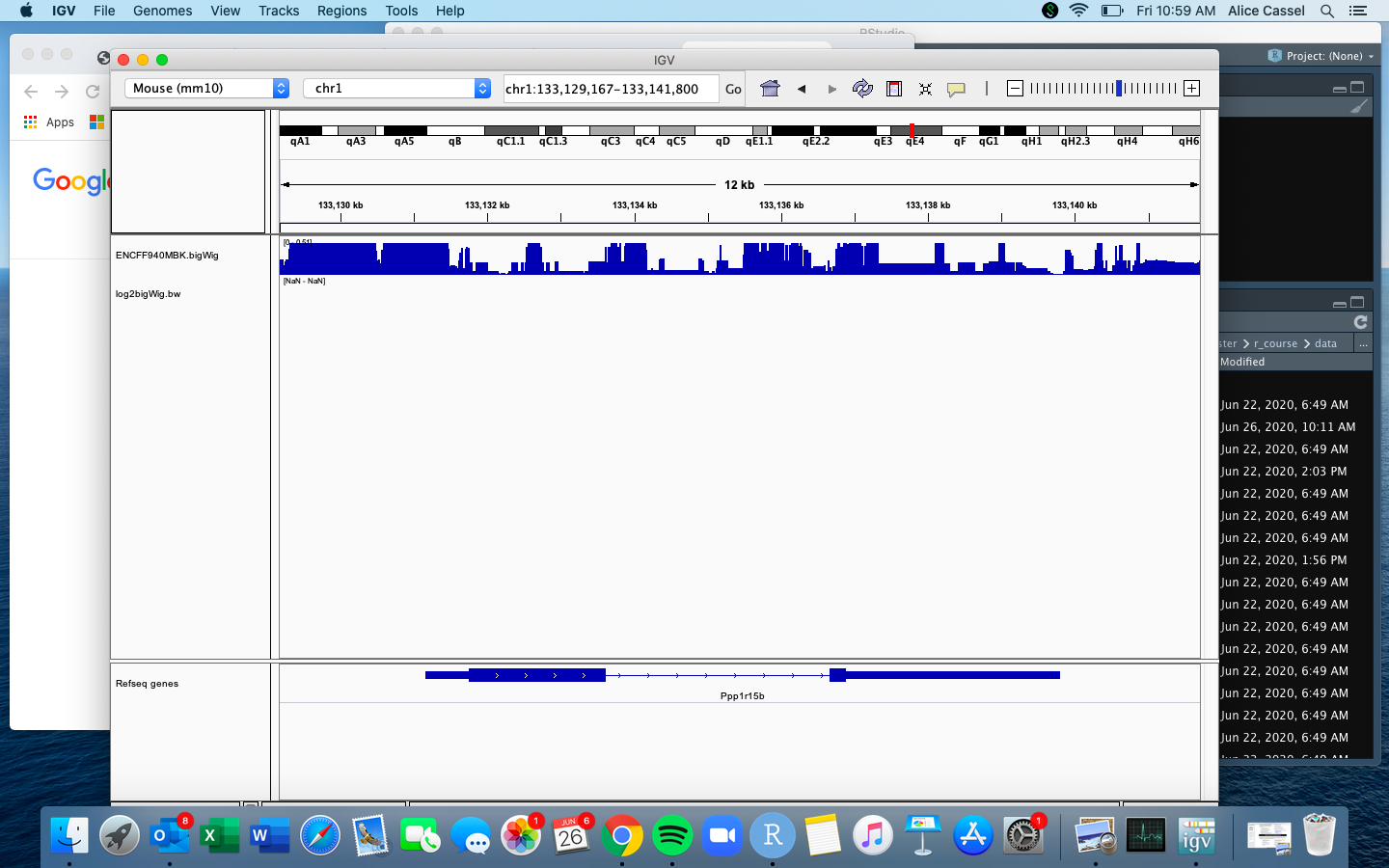
When I try to export the log2 transformed bigWig file to IGV, I get no visual peaks and the [NaN-NaN] message (see screenshot).
My code:
newBigWig <- myBigWig
newBigWig <- log2(newBigWig)
export.bw(newBigWig,"log2bigWig.bw")
The rest of my code from the exercise so far in case the problem is earlier (all has worked so far through making geom_area plot):
#Friday lecture: starting with slides 4
#genomic scores
suppressPackageStartupMessages(library(rtracklayer))
suppressPackageStartupMessages(library(GenomicRanges))
importRegion <- GRanges(seqnames="chr1",ranges=IRanges(133040000,133149400))
mySelection <- BigWigSelection(importRegion)
myBigWig <- import.bw("data/ENCFF940MBK.bigWig",selection=mySelection,as="RleList")
myBigWig
theMax <- max(myBigWig$chr1[133040000:133149400])
theMax
theMin <- min(myBigWig$chr1[133040000:133149400])
theMin
library(ggplot2)
TStartSite <- seq(133131166-500,133131166+500)
TSSdf <- as.data.frame(myBigWig$chr1[TStartSite])
TSSdf$position <- TStartSite
ggplot(TSSdf,aes(x=position,y=value))+geom_area()+theme_minimal()
Reactions are currently unavailable